RETINOIC ACID METABOLIC PATHWAY (PW:0001004) Description
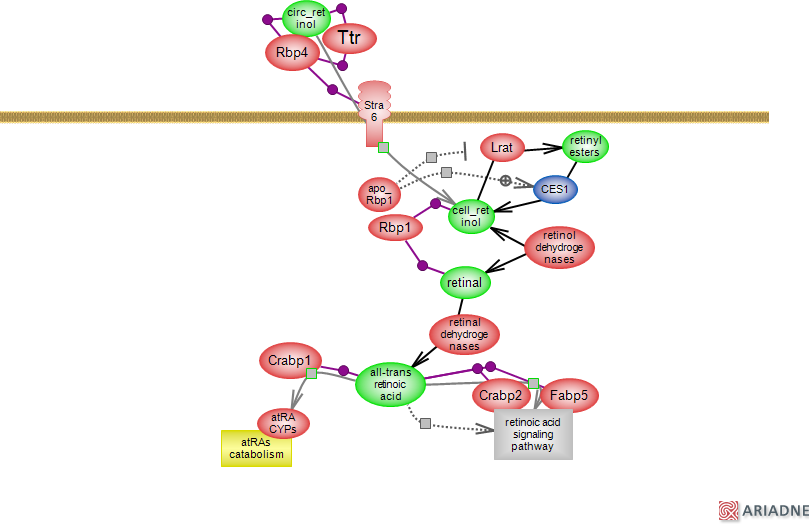
Animal species cannot synthesize retinol or vitamin A. They derive it from precursor carotenoids such as beta-carotene via retinal and they can store it as retinyl esters (REs) which they can hydrolyze (liver is the largest REs storage). Thus, both plant and animal dietary sources can provide vitamin A. Vitamin A metabolites such as retinal and retinoic acid play essential roles in vision and a range of cellular processes. More recently, a role for retinol in mitochondrial signaling where it may serve as a co-factor in redox sensing and energy homeostasis, has been postulated. The lipid retinol is subject to oxidation and as such is associated with carrier proteins such as Rbp4 in the circulation and Rbp1 inside cells. Rbp4 associates with transthyretin (Ttr) that also transports thyroxine (T4) and can act as a protease with important functions in the nervous system. The Ttr-Rbp4 association is stabilized by retinol but not by other retinoids. The ternary complex consists of a dimer of dimers Ttr tetramer with two Rbp4 molecules bound on either side of the dimers; in the complex, retinol also establishes contacts with Ttr (
PDB entry for the ternary complex ). Stra6 acts as a receptor for Rbp4 and as a transporter for retinol. Interestingly, Stra6 may also act as a signal transducer by inducing the Jak-Stat pathway. Inside the cell, retinol may be esterified to REs by the lecithin retinol acyltransferase Lrat in most tissues. REs are converted back to retinol by the action of retinyl ester hydrolase CES1. The human CES1 has a mouse but not a rat ortholog; several rat and mouse genes are potential REs hydrolases as well as a number of lipases (human CES1 gene shown). The ratio of apo- to holo-Rbp1 maintains a balanced flux of retinol to and from its REs storage. Apo-Rbp1 stimulates hydrolysis of REs while inhibiting retinol esterification. Alternatively, retinol may be reversibly converted to retinal by members of the short chain dehydrogenase/reductase family (SDR). In the retinal pigment epithelium, retinol is metabolized to cis-retinal to serve as the chromophore of the visual pigment. In other cells, retinal is the substrate for retinal dehydrogenases that irreversible convert it to all-trans retinoic acid (atRA) and other derivatives with atRA being the most potent molecule. atRA interacts with several proteins of the FABP family that exert distinct roles. Crabp2 and Fabp5 deliver atRA to particular nuclear receptors such as retinoid acid receptors (RARs) and a subset of peroxisome proliferator-activated receptors (PPAR), respectively. RARs form heterodimers with retinoid X receptors (RXR)s which partner with a number of other nuclear receptors, do not interact with atRA but favor 9-cisRA. Crabp1 on the other hand, relates to the processing and catabolism of retinoic acid. While the exact role of Crabp1 is still to be documented, its overexpression increases the rate of atRA catabolism. Several members of the cytochrome P450 are involved in the process. atRA signaling is important for development, cell proliferation, differentiation and death, immune and nervous systems function. Alterations in these signaling pathways has been associated with several conditions including cancer.
To see the ontology report for annotations, Gviewer and download, click here [click to see associated GO term GO:0042573 ] ...(less)
Pathway Diagram:
GO TO:
Genes in Pathway:
Symbol Object Name Evidence Notes Source PubMed Reference(s) RGD Reference(s) Position
G
Aldh1a1 aldehyde dehydrogenase 1 family, member A1
TAS IMP RGD
PMID:21621639 PMID:21138835 RGD:6484672 , RGD:6771354
NCBI chr 1:227,426,939...227,579,497
G
Aldh1a2 aldehyde dehydrogenase 1 family, member A2
TAS IEP IMP RGD
PMID:21621639 PMID:15950969 PMID:21138835 RGD:6484672 , RGD:6771322 , RGD:6771354
NCBI chr 8:80,758,641...80,837,891
G
Aldh1a3 aldehyde dehydrogenase 1 family, member A3
ISO IEP IMP RGD
PMID:21621639 PMID:15950969 PMID:21138835 RGD:6484672 , RGD:6771322 , RGD:6771354
NCBI chr 1:129,392,516...129,436,552
G
Ces1d carboxylesterase 1D
ISO RGD PMID:21621639 RGD:6484672
NCBI chr19:30,046,494...30,085,039
G
Crabp1 cellular retinoic acid binding protein 1
ISO RGD
PMID:21621639 RGD:6484672
NCBI chr 8:64,047,431...64,055,469
G
Crabp2 cellular retinoic acid binding protein 2
ISO IEP RGD
PMID:21621639 PMID:15950969 RGD:6484672 , RGD:6771322
NCBI chr 2:175,714,862...175,719,208
G
Cyp26a1 cytochrome P450, family 26, subfamily a, polypeptide 1
ISO RGD
PMID:21621639 RGD:6484672
NCBI chr 1:244,883,822...244,887,657
G
Cyp26b1 cytochrome P450, family 26, subfamily b, polypeptide 1
ISO IMP RGD
PMID:21621639 PMID:21138835 RGD:6484672 , RGD:6771354
NCBI chr 4:118,599,356...118,616,176
G
Cyp26c1 cytochrome P450, family 26, subfamily C, polypeptide 1
ISO RGD
PMID:21621639 RGD:6484672
NCBI chr 1:244,870,412...244,881,610
G
Dhrs9 dehydrogenase/reductase 9
ISO IEP IMP RGD
PMID:21621639 PMID:15950969 PMID:21138835 RGD:6484672 , RGD:6771322 , RGD:6771354
NCBI chr 3:74,553,357...74,579,535
G
Fabp5 fatty acid binding protein 5
ISO RGD
PMID:21621639 RGD:6484672
NCBI chr 2:93,672,451...93,676,186
G
Lrat lecithin retinol acyltransferase
ISO RGD
PMID:21621639 RGD:6484672
NCBI chr 2:170,562,148...170,571,212
G
Rbp1 retinol binding protein 1
ISO IEP RGD
PMID:21621639 PMID:15950969 RGD:6484672 , RGD:6771322
NCBI chr 8:107,904,589...107,926,109
G
Rbp4 retinol binding protein 4
ISO RGD
PMID:21782034 PMID:21621639 RGD:6484671 , RGD:6484672
NCBI chr 1:245,306,349...245,313,551
G
Rdh10 retinol dehydrogenase 10
ISO IMP RGD
PMID:21621639 PMID:21138835 RGD:6484672 , RGD:6771354
NCBI chr 5:3,186,671...3,215,572
G
Rdh16 retinol dehydrogenase 16
IMP RGD
PMID:21138835 RGD:6771354
NCBI chr 7:65,499,763...65,510,304
G
Rdh7 retinol dehydrogenase 7
TAS RGD
PMID:21621639 RGD:6484672
NCBI chr 7:63,676,016...63,689,375
G
Stra6 signaling receptor and transporter of retinol STRA6
ISO RGD
PMID:21782034 PMID:21621639 RGD:6484671 , RGD:6484672
NCBI chr 8:67,444,757...67,464,719
G
Ttr transthyretin
ISO RGD
PMID:21782034 RGD:6484671
NCBI chr18:12,216,684...12,225,972
Additional Elements in Pathway: (includes Gene Groups, Small Molecules, Other Pathways..etc.) Small Molecule circ_retinol circulating retinol Functional Class retinol dehydrogenases enzymes that catalyze the reversible conversion of retinol to retinal Small Molecule cell_retinol intracellular retinol Functional Class atRA CYPs the cytochrome P450 family 26 members involved in the catabolism of retinoic acid Functional Class retinal dehydrogenases the enzymes the catalyze the irreversible conversion of retinal to retinoic acid
Pathway Gene Annotations
Disease Annotations Associated with Genes in the retinoic acid metabolic pathway
Aldh1a1 autism spectrum disorder , Creutzfeldt-Jakob disease , Experimental Liver Cirrhosis , liver disease , melanoma , metabolic dysfunction-associated steatotic liver disease , Neoplastic Cell Transformation , oral squamous cell carcinoma , pancreatic cancer , Parkinsonism , renal cell carcinoma Aldh1a2 Bloom syndrome , colorectal cancer , Coronary Occlusion , Diaphragmatic Hernia 4 , DiGeorge syndrome , duodenal atresia , limb ischemia , myelomeningocele , osteoarthritis , Parkinsonism , Prostatic Neoplasms , Schistosomiasis Mansoni Aldh1a3 autism spectrum disorder , autistic disorder , chromosome 15q26-qter deletion syndrome , genetic disease , isolated microphthalmia 8 , melanoma , microphthalmia , Neurodevelopmental Disorders , Stomach Neoplasms , syndromic microphthalmia 9 Ces1d alcohol dependence , Brain Injuries , Chromosome 16q12 Duplication Syndrome , Cocaine-Related Disorders , colorectal cancer , COVID-19 , gastrointestinal system cancer , hepatocellular carcinoma , Hypercholesterolemia , inherited metabolic disorder , liver cancer , Lung Neoplasms , Monocyte Esterase Deficiency , obesity , opiate dependence , stomach cancer Crabp1 Bloom syndrome , colorectal cancer , esophagus squamous cell carcinoma , Keloid , Liver Neoplasms , renal cell carcinoma Crabp2 Charcot-Marie-Tooth disease type 2 , gastrointestinal stromal tumor , immunodeficiency 42 , methylmalonic acidemia , MHC class II deficiency , Ovarian Neoplasms , parathyroid carcinoma , Recurrence , severe congenital neutropenia 3 , severe congenital neutropenia 5 Cyp26a1 actinic keratosis , adenocarcinoma , Barrett's adenocarcinoma , Barrett's esophagus , Colonic Neoplasms , endometriosis , Esophageal Neoplasms , Experimental Liver Cirrhosis , familial adenomatous polyposis , spina bifida , Vitamin A Deficiency Cyp26b1 autism spectrum disorder , congenital disorder of glycosylation type IIb , Craniosynostosis Syndrome, Autosomal Recessive , dystonia , esophagus squamous cell carcinoma , genetic disease , hepatocellular carcinoma , Methylmalonyl-CoA Epimerase Deficiency , mouth disease , obesity , Radiohumeral Fusions with Other Skeletal and Craniofacial Anomalies Cyp26c1 Focal Facial Dermal Dysplasia , Focal Facial Dermal Dysplasia 4 , syndromic microphthalmia 5 Dhrs9 Bardet-Biedl syndrome , Birth Weight Fabp5 hepatocellular carcinoma , Myocardial Ischemia , prostate cancer , psoriasis Lrat breast ductal carcinoma , celiac disease , Chemical and Drug Induced Liver Injury , cone-rod dystrophy , Experimental Liver Cirrhosis , Eye Abnormalities , fundus dystrophy , genetic disease , hepatocellular carcinoma , Hereditary Eye Diseases , invasive ductal carcinoma , Leber congenital amaurosis , Leber congenital amaurosis 1 , Leber congenital amaurosis 14 , Leber hereditary optic neuropathy , retinitis pigmentosa , Vitamin A Deficiency Rbp1 Animal Mammary Neoplasms , basal cell carcinoma , carcinoma , Chemical and Drug Induced Liver Injury , Experimental Mammary Neoplasms , Fetal Growth Retardation , Leber congenital amaurosis , multiple myeloma , Stomach Neoplasms Rbp4 Acute Coronary Syndrome , acute pyelonephritis , Alcoholic Liver Diseases , Animal Mammary Neoplasms , Anophthalmia , anterior segment dysgenesis , aortic disease , Arteriovenous Fistula , atherosclerosis , Breast Neoplasms , carcinoma , Cardiomegaly , Carotid Atherosclerosis , cerebral infarction , chronic kidney disease , coloboma , colon adenoma , congestive heart failure , coronary artery disease , Coronary Disease , diabetic angiopathy , Diabetic Nephropathies , diabetic retinopathy , dilated cardiomyopathy , Dyslipidemias , essential hypertension , Experimental Liver Cirrhosis , Experimental Mammary Neoplasms , Eye Abnormalities , familial hypercholesterolemia , familial temporal lobe epilepsy 1 , fundus dystrophy , gallbladder cancer , genetic disease , gestational diabetes , hyperinsulinism , hypertension , Hypertriglyceridemia , Insulin Resistance , Jaundice , keratomalacia , kidney disease , liver cirrhosis , Metabolic Syndrome , microphthalmia , Microphthalmia/Coloboma 10 , myocardial infarction , Necrosis , obesity , Pediatric Obesity , polycystic ovary syndrome , pre-eclampsia , Pregnancy-Induced Hypertension , Prehypertension , proliferative diabetic retinopathy , psoriasis , Retinal Dystrophy, Iris Coloboma, and Comedogenic Acne Syndrome , severe pre-eclampsia , Spontaneous Abortions , Stable Angina , Stomach Neoplasms , systemic scleroderma , type 2 diabetes mellitus , Vision Disorders , Vitamin A Deficiency Rdh10 Craniofacial Abnormalities , Experimental Liver Cirrhosis , Fetal Death Rdh16 cataract 15 multiple types , Charcot-Marie-Tooth disease axonal type 2U , INTERSTITIAL LUNG AND LIVER DISEASE , paraplegia Stra6 Anophthalmia , Bardet-Biedl syndrome , Bloom syndrome , Brugada syndrome 8 , Cardiovascular Abnormalities , colorectal adenocarcinoma , colorectal cancer , congenital heart disease , Craniofacial Abnormalities , diabetic retinopathy , Diaphragmatic Hernia , exfoliation syndrome , Experimental Diabetes Mellitus , genetic disease , intellectual disability , Isolated Microphthalmia with Coloboma , lung disease , microphthalmia , PAPA syndrome , persistent fetal circulation syndrome , schizophrenia , syndromic microphthalmia 5 , syndromic microphthalmia 9 Ttr acute kidney failure , adenosquamous lung carcinoma , Alzheimer's disease , Amyloid Neuropathies , amyloidosis , anemia , arrhythmogenic right ventricular dysplasia 10 , arrhythmogenic right ventricular dysplasia 11 , bone marrow disease , brain ischemia , Brugada syndrome , cardiac amyloidosis , Cardiac Conduction Defect , cardiomyopathy , carpal tunnel syndrome , carpal tunnel syndrome 1 , Charcot-Marie-Tooth disease , Chemical and Drug Induced Liver Injury , cholangiocarcinoma , chromosome 18q deletion syndrome , colorectal adenoma , colorectal cancer , congestive heart failure , Drug Eruptions , Drug-Related Side Effects and Adverse Reactions , dystransthyretinemic hyperthyroxinemia , Familial Amyloid Polyneuropathies , Familial Amyloidosis , familial hypertrophic cardiomyopathy , Heart Block , hepatocellular carcinoma , hypertrophic cardiomyopathy , hypertrophic cardiomyopathy 1 , Hypothermia , intellectual disability , lung adenocarcinoma , lung cancer , lung large cell carcinoma , Lung Neoplasms , lung non-small cell carcinoma , lung small cell carcinoma , Lymphatic Metastasis , Neoplasm Metastasis , oral squamous cell carcinoma , pancytopenia , stomach cancer , Sudden Unexpected Nocturnal Death Syndrome , transthyretin amyloidosis
actinic keratosis Cyp26a1 Acute Coronary Syndrome Rbp4 acute kidney failure Ttr acute pyelonephritis Rbp4 adenocarcinoma Cyp26a1 adenosquamous lung carcinoma Ttr alcohol dependence Ces1d Alcoholic Liver Diseases Rbp4 Alzheimer's disease Ttr Amyloid Neuropathies Ttr amyloidosis Ttr anemia Ttr Animal Mammary Neoplasms Rbp1 , Rbp4 Anophthalmia Rbp4 , Stra6 anterior segment dysgenesis Rbp4 aortic disease Rbp4 arrhythmogenic right ventricular dysplasia 10 Ttr arrhythmogenic right ventricular dysplasia 11 Ttr Arteriovenous Fistula Rbp4 atherosclerosis Rbp4 autism spectrum disorder Aldh1a1 , Aldh1a3 , Cyp26b1 autistic disorder Aldh1a3 Bardet-Biedl syndrome Dhrs9 , Stra6 Barrett's adenocarcinoma Cyp26a1 Barrett's esophagus Cyp26a1 basal cell carcinoma Rbp1 Birth Weight Dhrs9 Bloom syndrome Aldh1a2 , Crabp1 , Stra6 bone marrow disease Ttr Brain Injuries Ces1d brain ischemia Ttr breast ductal carcinoma Lrat Breast Neoplasms Rbp4 Brugada syndrome Ttr Brugada syndrome 8 Stra6 carcinoma Rbp1 , Rbp4 cardiac amyloidosis Ttr Cardiac Conduction Defect Ttr Cardiomegaly Rbp4 cardiomyopathy Ttr Cardiovascular Abnormalities Stra6 Carotid Atherosclerosis Rbp4 carpal tunnel syndrome Ttr carpal tunnel syndrome 1 Ttr cataract 15 multiple types Rdh16 celiac disease Lrat cerebral infarction Rbp4 Charcot-Marie-Tooth disease Ttr Charcot-Marie-Tooth disease axonal type 2U Rdh16 Charcot-Marie-Tooth disease type 2 Crabp2 Chemical and Drug Induced Liver Injury Lrat , Rbp1 , Ttr cholangiocarcinoma Ttr chromosome 15q26-qter deletion syndrome Aldh1a3 Chromosome 16q12 Duplication Syndrome Ces1d chromosome 18q deletion syndrome Ttr chronic kidney disease Rbp4 Cocaine-Related Disorders Ces1d coloboma Rbp4 colon adenoma Rbp4 Colonic Neoplasms Cyp26a1 colorectal adenocarcinoma Stra6 colorectal adenoma Ttr colorectal cancer Aldh1a2 , Ces1d , Crabp1 , Stra6 , Ttr cone-rod dystrophy Lrat congenital disorder of glycosylation type IIb Cyp26b1 congenital heart disease Stra6 congestive heart failure Rbp4 , Ttr coronary artery disease Rbp4 Coronary Disease Rbp4 Coronary Occlusion Aldh1a2 COVID-19 Ces1d Craniofacial Abnormalities Rdh10 , Stra6 Craniosynostosis Syndrome, Autosomal Recessive Cyp26b1 Creutzfeldt-Jakob disease Aldh1a1 diabetic angiopathy Rbp4 Diabetic Nephropathies Rbp4 diabetic retinopathy Rbp4 , Stra6 Diaphragmatic Hernia Stra6 Diaphragmatic Hernia 4 Aldh1a2 DiGeorge syndrome Aldh1a2 dilated cardiomyopathy Rbp4 Drug Eruptions Ttr Drug-Related Side Effects and Adverse Reactions Ttr duodenal atresia Aldh1a2 Dyslipidemias Rbp4 dystonia Cyp26b1 dystransthyretinemic hyperthyroxinemia Ttr endometriosis Cyp26a1 Esophageal Neoplasms Cyp26a1 esophagus squamous cell carcinoma Crabp1 , Cyp26b1 essential hypertension Rbp4 exfoliation syndrome Stra6 Experimental Diabetes Mellitus Stra6 Experimental Liver Cirrhosis Aldh1a1 , Cyp26a1 , Lrat , Rbp4 , Rdh10 Experimental Mammary Neoplasms Rbp1 , Rbp4 Eye Abnormalities Lrat , Rbp4 familial adenomatous polyposis Cyp26a1 Familial Amyloid Polyneuropathies Ttr Familial Amyloidosis Ttr familial hypercholesterolemia Rbp4 familial hypertrophic cardiomyopathy Ttr familial temporal lobe epilepsy 1 Rbp4 Fetal Death Rdh10 Fetal Growth Retardation Rbp1 Focal Facial Dermal Dysplasia Cyp26c1 Focal Facial Dermal Dysplasia 4 Cyp26c1 fundus dystrophy Lrat , Rbp4 gallbladder cancer Rbp4 gastrointestinal stromal tumor Crabp2 gastrointestinal system cancer Ces1d genetic disease Aldh1a3 , Cyp26b1 , Lrat , Rbp4 , Stra6 gestational diabetes Rbp4 Heart Block Ttr hepatocellular carcinoma Ces1d , Cyp26b1 , Fabp5 , Lrat , Ttr Hereditary Eye Diseases Lrat Hypercholesterolemia Ces1d hyperinsulinism Rbp4 hypertension Rbp4 Hypertriglyceridemia Rbp4 hypertrophic cardiomyopathy Ttr hypertrophic cardiomyopathy 1 Ttr Hypothermia Ttr immunodeficiency 42 Crabp2 inherited metabolic disorder Ces1d Insulin Resistance Rbp4 intellectual disability Stra6 , Ttr INTERSTITIAL LUNG AND LIVER DISEASE Rdh16 invasive ductal carcinoma Lrat isolated microphthalmia 8 Aldh1a3 Isolated Microphthalmia with Coloboma Stra6 Jaundice Rbp4 Keloid Crabp1 keratomalacia Rbp4 kidney disease Rbp4 Leber congenital amaurosis Lrat , Rbp1 Leber congenital amaurosis 1 Lrat Leber congenital amaurosis 14 Lrat Leber hereditary optic neuropathy Lrat limb ischemia Aldh1a2 liver cancer Ces1d liver cirrhosis Rbp4 liver disease Aldh1a1 Liver Neoplasms Crabp1 lung adenocarcinoma Ttr lung cancer Ttr lung disease Stra6 lung large cell carcinoma Ttr Lung Neoplasms Ces1d , Ttr lung non-small cell carcinoma Ttr lung small cell carcinoma Ttr Lymphatic Metastasis Ttr melanoma Aldh1a1 , Aldh1a3 metabolic dysfunction-associated steatotic liver disease Aldh1a1 Metabolic Syndrome Rbp4 methylmalonic acidemia Crabp2 Methylmalonyl-CoA Epimerase Deficiency Cyp26b1 MHC class II deficiency Crabp2 microphthalmia Aldh1a3 , Rbp4 , Stra6 Microphthalmia/Coloboma 10 Rbp4 Monocyte Esterase Deficiency Ces1d mouth disease Cyp26b1 multiple myeloma Rbp1 myelomeningocele Aldh1a2 myocardial infarction Rbp4 Myocardial Ischemia Fabp5 Necrosis Rbp4 Neoplasm Metastasis Ttr Neoplastic Cell Transformation Aldh1a1 Neurodevelopmental Disorders Aldh1a3 obesity Ces1d , Cyp26b1 , Rbp4 opiate dependence Ces1d oral squamous cell carcinoma Aldh1a1 , Ttr osteoarthritis Aldh1a2 Ovarian Neoplasms Crabp2 pancreatic cancer Aldh1a1 pancytopenia Ttr PAPA syndrome Stra6 paraplegia Rdh16 parathyroid carcinoma Crabp2 Parkinsonism Aldh1a1 , Aldh1a2 Pediatric Obesity Rbp4 persistent fetal circulation syndrome Stra6 polycystic ovary syndrome Rbp4 pre-eclampsia Rbp4 Pregnancy-Induced Hypertension Rbp4 Prehypertension Rbp4 proliferative diabetic retinopathy Rbp4 prostate cancer Fabp5 Prostatic Neoplasms Aldh1a2 psoriasis Fabp5 , Rbp4 Radiohumeral Fusions with Other Skeletal and Craniofacial Anomalies Cyp26b1 Recurrence Crabp2 renal cell carcinoma Aldh1a1 , Crabp1 Retinal Dystrophy, Iris Coloboma, and Comedogenic Acne Syndrome Rbp4 retinitis pigmentosa Lrat Schistosomiasis Mansoni Aldh1a2 schizophrenia Stra6 severe congenital neutropenia 3 Crabp2 severe congenital neutropenia 5 Crabp2 severe pre-eclampsia Rbp4 spina bifida Cyp26a1 Spontaneous Abortions Rbp4 Stable Angina Rbp4 stomach cancer Ces1d , Ttr Stomach Neoplasms Aldh1a3 , Rbp1 , Rbp4 Sudden Unexpected Nocturnal Death Syndrome Ttr syndromic microphthalmia 5 Cyp26c1 , Stra6 syndromic microphthalmia 9 Aldh1a3 , Stra6 systemic scleroderma Rbp4 transthyretin amyloidosis Ttr type 2 diabetes mellitus Rbp4 Vision Disorders Rbp4 Vitamin A Deficiency Cyp26a1 , Lrat , Rbp4
Pathway Annotations Associated with Genes in the retinoic acid metabolic pathway
Aldh1a1 arginine and proline metabolic pathway , ascorbate and aldarate metabolic pathway , beta-alanine metabolic pathway , cyclophosphamide pharmacodynamics pathway , cyclophosphamide pharmacokinetics pathway , fatty acid metabolic pathway , gluconeogenesis pathway , glycerolipid metabolic pathway , glycolysis pathway , glycolysis/gluconeogenesis pathway , histidine metabolic pathway , ifosfamide pharmacodynamics pathway , ifosfamide pharmacokinetics pathway , lysine degradation pathway , neviparine pharmacokinetics pathway , pentose and glucuronate interconversion pathway , propanoate metabolic pathway , pyruvate metabolic pathway , retinoic acid metabolic pathway , retinol metabolic pathway , tryptophan metabolic pathway , valine, leucine and isoleucine degradation pathway , vitamin A deficiency pathway Aldh1a2 retinoic acid metabolic pathway , retinol metabolic pathway , vitamin A deficiency pathway Aldh1a3 beta-alanine metabolic pathway , gluconeogenesis pathway , glycolysis pathway , glycolysis/gluconeogenesis pathway , histidine metabolic pathway , phase I biotransformation pathway via cytochrome P450 , phenylalanine metabolic pathway , retinoic acid metabolic pathway , tyrosine metabolic pathway Ces1d capecitabine pharmacodynamics pathway , capecitabine pharmacokinetics pathway , heroin pharmacodynamics pathway , heroin pharmacokinetics pathway , irinotecan pharmacodynamics pathway , irinotecan pharmacokinetics pathway , mycophenolic acid pharmacokinetics pathway , retinoic acid metabolic pathway Crabp1 retinoic acid metabolic pathway Crabp2 retinoic acid metabolic pathway Cyp26a1 retinoic acid metabolic pathway , retinol metabolic pathway , vitamin A deficiency pathway Cyp26b1 retinoic acid metabolic pathway , retinol metabolic pathway Cyp26c1 retinoic acid metabolic pathway , retinol metabolic pathway Dhrs9 retinoic acid metabolic pathway , retinol metabolic pathway , vitamin A deficiency pathway Fabp5 eicosanoid signaling pathway via peroxisome proliferator-activated receptor gamma , retinoic acid metabolic pathway Lrat altered retinoid cycle metabolic pathway , retinitis pigmentosa pathway , retinoic acid metabolic pathway , retinoid cycle metabolic pathway , retinol metabolic pathway , vitamin A deficiency pathway Rbp1 retinoic acid metabolic pathway , retinoid cycle metabolic pathway Rbp4 retinoic acid metabolic pathway Rdh10 retinoic acid metabolic pathway , retinoid cycle metabolic pathway , retinol metabolic pathway Rdh16 retinoic acid metabolic pathway , retinol metabolic pathway , vitamin A deficiency pathway Rdh7 retinoic acid metabolic pathway , retinol metabolic pathway Stra6 retinoic acid metabolic pathway Ttr forkhead class A signaling pathway , retinoic acid metabolic pathway
altered retinoid cycle metabolic pathway Lrat arginine and proline metabolic pathway Aldh1a1 ascorbate and aldarate metabolic pathway Aldh1a1 beta-alanine metabolic pathway Aldh1a1 , Aldh1a3 capecitabine pharmacodynamics pathway Ces1d capecitabine pharmacokinetics pathway Ces1d cyclophosphamide pharmacodynamics pathway Aldh1a1 cyclophosphamide pharmacokinetics pathway Aldh1a1 eicosanoid signaling pathway via peroxisome proliferator-activated receptor gamma Fabp5 fatty acid metabolic pathway Aldh1a1 forkhead class A signaling pathway Ttr gluconeogenesis pathway Aldh1a1 , Aldh1a3 glycerolipid metabolic pathway Aldh1a1 glycolysis pathway Aldh1a1 , Aldh1a3 glycolysis/gluconeogenesis pathway Aldh1a1 , Aldh1a3 heroin pharmacodynamics pathway Ces1d heroin pharmacokinetics pathway Ces1d histidine metabolic pathway Aldh1a1 , Aldh1a3 ifosfamide pharmacodynamics pathway Aldh1a1 ifosfamide pharmacokinetics pathway Aldh1a1 irinotecan pharmacodynamics pathway Ces1d irinotecan pharmacokinetics pathway Ces1d lysine degradation pathway Aldh1a1 mycophenolic acid pharmacokinetics pathway Ces1d neviparine pharmacokinetics pathway Aldh1a1 pentose and glucuronate interconversion pathway Aldh1a1 phase I biotransformation pathway via cytochrome P450 Aldh1a3 phenylalanine metabolic pathway Aldh1a3 propanoate metabolic pathway Aldh1a1 pyruvate metabolic pathway Aldh1a1 retinitis pigmentosa pathway Lrat retinoic acid metabolic pathway Aldh1a1 , Aldh1a2 , Aldh1a3 , Ces1d , Crabp1 , Crabp2 , Cyp26a1 , Cyp26b1 , Cyp26c1 , Dhrs9 , Fabp5 , Lrat , Rbp1 , Rbp4 , Rdh10 , Rdh16 , Rdh7 , Stra6 , Ttr retinoid cycle metabolic pathway Lrat , Rbp1 , Rdh10 retinol metabolic pathway Aldh1a1 , Aldh1a2 , Cyp26a1 , Cyp26b1 , Cyp26c1 , Dhrs9 , Lrat , Rdh10 , Rdh16 , Rdh7 tryptophan metabolic pathway Aldh1a1 tyrosine metabolic pathway Aldh1a3 valine, leucine and isoleucine degradation pathway Aldh1a1 vitamin A deficiency pathway Aldh1a1 , Aldh1a2 , Cyp26a1 , Dhrs9 , Lrat , Rdh16
References Associated with the retinoic acid metabolic pathway:
D'Ambrosio DN, etal., Nutrients. 2011 Jan;3(1):63-103.
Berry DC and Noy N, Biochim Biophys Acta. 2012 Jan;1821(1):168-76. Epub 2011 Jul 12.
Napoli JL Biochim Biophys Acta. 2012 Jan;1821(1):152-67. Epub 2011 May 19.
Germain P, etal., Pharmacol Rev. 2006 Dec;58(4):712-25.
Germain P, etal., Pharmacol Rev. 2006 Dec;58(4):760-72.
Liz MA, etal., IUBMB Life. 2010 Jun;62(6):429-35.
Schreiber R, etal., Biochim Biophys Acta. 2012 Jan;1821(1):113-23. Epub 2011 May 8.
Harrison EH Biochim Biophys Acta. 2012 Jan;1821(1):70-7. Epub 2011 Jun 12.
Kane MA Biochim Biophys Acta. 2012 Jan;1821(1):10-20. Epub 2011 Oct 1.
Theodosiou M, etal., Cell Mol Life Sci. 2010 May;67(9):1423-45. Epub 2010 Feb 7.
Shirakami Y, etal., Biochim Biophys Acta. 2012 Jan;1821(1):124-36. Epub 2011 Jul 1.
Dawson MI and Xia Z, Biochim Biophys Acta. 2012 Jan;1821(1):21-56. Epub 2011 Oct 1.
Ontology Path Diagram:
Import into Pathway Studio: